G. Matthew Fricke
 I am a Research Associate Professor of Computer Science at the
University of New Mexico, where I study computation in complex decentralised systems like supercomputing clusters, robot swarms,
social insects, and immune systems. I also have an interest in the ethics of artificial systems and complexity measures as biosignatures.
The robotics and computational biology research is funded by the Moses Biological Computation Lab.
Through the Center for Advanced Research Computing I collaborate with scientists from a variety of disciplines
to scale up their computations to supercomputers. I enjoy teaching whenever I can and mentoring students.
I am a Research Associate Professor of Computer Science at the
University of New Mexico, where I study computation in complex decentralised systems like supercomputing clusters, robot swarms,
social insects, and immune systems. I also have an interest in the ethics of artificial systems and complexity measures as biosignatures.
The robotics and computational biology research is funded by the Moses Biological Computation Lab.
Through the Center for Advanced Research Computing I collaborate with scientists from a variety of disciplines
to scale up their computations to supercomputers. I enjoy teaching whenever I can and mentoring students.
I earned a BA in Anthropology (Archaeology) from Appalachian State University; and a BS in Mathematics, and MS (Artificial Intelligence) and PhD degrees in Computer Science from the University of New Mexico. Through my consulting company, Go Figure Software, I've supported researchers in the UNM Physics and Los Alamos National Labs with custom scientific software. Archa{e}oImaging is a project to produce multi-spectral maps of archaeological sites.
My wife, Suzanne, and I have four sons: Henry, Leo, Owen, and Tristan.
Suzanne is an art historian (CV) who taught courses ranging from the Palaeolithic to the post-modern at the University of New Mexico, the Institute of American Indian Arts, and the Santa Fe Institute for Art and Design. She owns Gallery Hózhó, a fine arts gallery for New Mexico artists in Albuquerque. She is also an author and curator, with publications including As We See It, Indigenous Futurisms: Transcending Past, Present, Future, and Indigenous Futurisms in the Hyperpresent Now, and she is a frequent contributor to First American Arts Magazine. Curated shows include Octopus Dreams and Future Imaginaries: Indigenous Art, Fashion, Technology. She earned her undergraduate degree from Mills College, her master's degree in Italian Renaissance art from the University of Chicago, and her PhD from UNM. Her dissertation, Institutionalising Taste: Kenneth Milton Chapman, the Indian Arts Fund, and the Growth of Fine Art Pueblo Pottery, examines the emergence of Pueblo pottery as fine art.
I was born and raised in Shrewsbury, UK, but have lived in Mount Airy, North Carolina and Albuquerque, New Mexico for most of my life. I have been interested in computers since playing with a BBC Computer when I was nine years old. I came to New Mexico for Harvard Ayers' southwest field school and stayed to work for the Park Service at Petroglyph National Monument.Office Schedule
Teaching
Courses and workshops
Courses
University teaching
Click to collapse
CS491/591: High Performance Computing
CS533: Experimental Methods in Computer Science
CS591: Programming Swarm Robots
CS523: Complex Adaptive Systems
CS261: Mathematical Foundations of Computer Science
CS151: Computer Programming Fundamentals with Python
CS151: Computer Programming Fundamentals with C++
HPC Workshops
CARC training materials
Click to collapse
Beginner Introduction: Basic Linux and Interactive SLURM
Intermediate Introduction: Batch SLURM with FORTRAN
Intermediate Introduction: Batch SLURM with Python
Intermediate Intro to SLURM
Intermediate Intro to SLURM with NAMD
SLURM and Gaussian (1 hr version)
Intermediate SLURM and StarCCM
PBS, Modules, GNU Parallel, Conda, and MPI
Parallel Processing with MPI and SLURM (50 min)
CS4/542: Introduction to Parallel Processing
CS523: Complex Adaptive Systems
CS/Math 471: Introduction to Scientific Computing
Chemistry 525: Structural Biology – Crystallography and CARC
Biology 4/519: Biodiversity Informatics
Quantum Computing for Quantum Chemistry
Global Climate Change
Computational Methods for Geoscience
Radio Astronomy and CASA
Scientific Computing Lecture (CS471, 2018)
Computational Fluid Dynamics
Quantum Computing
Bits and Pieces
Older course materials
Click to expand
Teaching materials index for search engines and crawlers
Direct links to courses, workshops, and teaching PDFs are listed here as plain HTML anchors so crawlers can discover them without relying on JavaScript or expanded panels.
- CS491/591: High Performance Computing (Web · Course — Course page)
- CS533: Experimental Methods in Computer Science (PDF · Course — Syllabus)
- CS591: Programming Swarm Robots (PDF · Course — Syllabus)
- CS523: Complex Adaptive Systems (Web · Course — Course page)
- CS261: Mathematical Foundations of Computer Science (Web · Course — Course page)
- CS151: Computer Programming Fundamentals with Python (PDF · Course — Syllabus)
- CS151: Computer Programming Fundamentals with C++ (Web · Course — Course page)
- Beginner Introduction: Basic Linux and Interactive SLURM (PDF · Workshop — Linux basics and interactive jobs)
- Intermediate Introduction: Batch SLURM with FORTRAN (PDF · Workshop — Batch jobs with FORTRAN examples)
- Intermediate Introduction: Batch SLURM with Python (PDF · Workshop — Batch jobs with Python examples)
- Intermediate Intro to SLURM (PDF · Workshop — Core SLURM concepts)
- Intermediate Intro to SLURM with NAMD (PDF · Workshop — MPI style workloads)
- SLURM and Gaussian (1 hr version) (PDF · Workshop — Short format workshop)
- Intermediate SLURM and StarCCM (PDF · Workshop — Application focused)
- PBS, Modules, GNU Parallel, Conda, and MPI (PDF · Workshop — Mixed tooling overview)
- Parallel Processing with MPI and SLURM (50 min) (PDF · Workshop — Concise MPI and scheduler introduction)
- CS4/542: Introduction to Parallel Processing (PDF · Workshop — Parallel fundamentals)
- CS523: Complex Adaptive Systems (Web · Workshop — Workshop notes)
- CS/Math 471: Introduction to Scientific Computing (PDF · Workshop — Scientific computing introduction)
- Chemistry 525: Structural Biology – Crystallography and CARC (PDF · Workshop — Crystallography and HPC workflow)
- Biology 4/519: Biodiversity Informatics (PDF · Workshop — Data and compute for biodiversity)
- Quantum Computing for Quantum Chemistry (PDF · Workshop — Chemistry oriented quantum computing)
- Global Climate Change (PDF · Workshop — Computational perspective)
- Computational Methods for Geoscience (PDF · Workshop — Methods and examples)
- Radio Astronomy and CASA (PDF · Workshop — CASA oriented HPC use)
- Computational Fluid Dynamics (PDF · Workshop — CFD focused workshop)
- Quantum Computing (PDF · Workshop — General quantum computing workshop)
- CARC Annual Meeting: Machine Learning (PDF · Workshop — Annual meeting session)
- ESCAPE Workshop with JupyterHub (PDF · Workshop — Notebook based workflows)
- CARC Workshop v1.5 (PDF · Workshop — Older overview)
- CARC Workshop v1.1 (PDF · Workshop — Older overview)
- CARC Workshop v1.4 (PDF · Workshop — Older overview)
- CS152: Computer Programming Fundamentals with Java (PDF · Web — OOP material)
- CS201: Discrete Mathematics (PDF — Lecture notes)
- CS251: Intermediate Programming with Java (Web — Course page)
- CS523: Complex Adaptive Systems (Spring 2013) (Web — Archived course page)
- Q-bio (Summer 2018) (PDF — Short course)
- Scientific Computing (HPC Lecture) (PDF — Standalone lecture)
- Coding for Beginners (Web — Intro materials)
Conferences
Online Chair (2024–26) for the IEEE Symposium on High-Performance Interconnects (HOTI), an annual conference established in 1993 that brings together researchers and practitioners in high-performance computing and networking.
Publications
My research explores how distributed systems acquire, process, and act on information, from immune cells and ant colonies to robot swarms, autonomous drones, infrastructure monitoring, climate systems, volcanology, and astrobiology.
PDF index for search engines and crawlers
Direct links to publication PDFs are listed here as plain HTML anchors so crawlers can discover them without relying on JavaScript filters.
- Automated uncrewed aerial system data acquisition for rebar layout inspection in the construction industry — Automated uncrewed aerial system data acquisition for rebar layout inspection in the construction industry (Journal · 2026)
- Development of a camera-laser-IMU system for non-contact transverse displacement estimation using an adaptive extended Kalman filter — Development of a camera-laser-IMU system for non-contact transverse displacement estimation using an adaptive extended Kalman filter (Journal · 2026)
- Bigger is faster in the adaptive immune response. — Bigger is faster in the adaptive immune response. (Journal · 2025)
- Space-Time Causal Discovery in Earth System Science: A Local Stencil Learning Approach. — Space-Time Causal Discovery in Earth System Science: A Local Stencil Learning Approach. (Journal · 2025)
- Social Network Analysis of Teams in Engineering and Computer Science Courses — Social Network Analysis of Teams in Engineering and Computer Science Courses (Conf. · 2025)
- More is Faster: Why Population Size Matters in Biological Search. — More is Faster: Why Population Size Matters in Biological Search. (Journal · 2024)
- CO2 emissions during the 2023 Litli Hrútur eruption in Reykjanes, Iceland: 13C tracks magma degassing. — CO2 emissions during the 2023 Litli Hrútur eruption in Reykjanes, Iceland: 13C tracks magma degassing. (Journal · 2024)
- Drone CO₂ measurements during the Tajogaite volcanic eruption. — Drone CO₂ measurements during the Tajogaite volcanic eruption. (Journal · 2024)
- Sensor Equipped UAS for Non-Contact Bridge Inspections: Field Application. — Sensor Equipped UAS for Non-Contact Bridge Inspections: Field Application. (Journal · 2023)
- Aerial Survey Robotics in Extreme Environments: Mapping Volcanic CO2 Emissions With Flocking UAVs. — Aerial Survey Robotics in Extreme Environments: Mapping Volcanic CO2 Emissions With Flocking UAVs. (Journal · 2022)
- The Grayness of the Origin of Life. — The Grayness of the Origin of Life. (Journal · 2021)
- Machine learning feature analysis illuminates disparity between E3SM climate models and observed climate change. — Machine learning feature analysis illuminates disparity between E3SM climate models and observed climate change. (Journal · 2021)
- Aerial strategies advance volcanic gas measurements at inaccessible, strongly degassing volcanoes. — Aerial strategies advance volcanic gas measurements at inaccessible, strongly degassing volcanoes. (Journal · 2020)
- Swarm Foraging Review: Closing the Gap Between Proof and Practice. — Swarm Foraging Review: Closing the Gap Between Proof and Practice. (Journal · 2020)
- Variability in the Analysis of a Single Neuroimaging Dataset by Many Teams. — Variability in the Analysis of a Single Neuroimaging Dataset by Many Teams. (Journal · 2020)
- Modeling T Cell Motion in Tissues During Immune Responses. — Modeling T Cell Motion in Tissues During Immune Responses. (Journal · 2019)
- Quantitative Measurement of Naïve T cell Association with Dendritic Cells, FRCs, and Blood Vessels in Lymph Nodes. — Quantitative Measurement of Naïve T cell Association with Dendritic Cells, FRCs, and Blood Vessels in Lymph Nodes. (Journal · 2018)
- ROCK regulates the intermittent mode of interstitial T cell migration in inflamed lungs. — ROCK regulates the intermittent mode of interstitial T cell migration in inflamed lungs. (Journal · 2017)
- Persistence and Adaptation in Immunity: T Cells Balance the Extent and Thoroughness of Search. — Persistence and Adaptation in Immunity: T Cells Balance the Extent and Thoroughness of Search. (Journal · 2016)
- Immune-inspired search strategies for robot swarms. — Immune-inspired search strategies for robot swarms. (Journal · 2016)
- Quantifying the Effect of Colony Size and Food Distribution on Harvester Ant Foraging. — Quantifying the Effect of Colony Size and Food Distribution on Harvester Ant Foraging. (Journal · 2012)
- GetBonNie for building, analyzing, and sharing rule-based models. — GetBonNie for building, analyzing, and sharing rule-based models. (Journal · 2009)
- Receptor aggregation by intermembrane interactions: A Monte Carlo study. — Receptor aggregation by intermembrane interactions: A Monte Carlo study. (Journal · 2006)
- Navigating the Edge: UAS Boundary Tracing for Efficient Volcanic Plume Monitoring. — Navigating the Edge: UAS Boundary Tracing for Efficient Volcanic Plume Monitoring. (Conf. · 2024)
- A Bio-Inspired Transportation Network for Scalable Swarm Foraging — A Bio-Inspired Transportation Network for Scalable Swarm Foraging (Conf. · 2020)
- LoCUS: A loss-tolerant volcano survey algorithm — LoCUS: A loss-tolerant volcano survey algorithm (Conf. · 2020)
- Ignorance is Not Bliss: An Analysis of Central-Place Foraging Algorithms. — Ignorance is Not Bliss: An Analysis of Central-Place Foraging Algorithms. (Conf. · 2019)
- Ignorance is Not Bliss: An Analysis of Central-Place Foraging Algorithms. — Presentation PDF (Conf. · 2019)
- Comparing physical and simulated performance of a deterministic and a bio-inspired stochastic foraging strategy for robot swarms — Comparing physical and simulated performance of a deterministic and a bio-inspired stochastic foraging strategy for robot swarms (Conf. · 2019)
- Brief Announcement: On Site Fidelity and the Price of Ignorance in Swarm Robotic Central Place Foraging Algorithms. — Brief Announcement: On Site Fidelity and the Price of Ignorance in Swarm Robotic Central Place Foraging Algorithms. (Conf. · 2019)
- A Distributed Deterministic Spiral Search Algorithm for Swarms. — A Distributed Deterministic Spiral Search Algorithm for Swarms. (Conf. · 2016)
- Distinguishing Adaptive Search From Random Search in Robots and T cells. — Distinguishing Adaptive Search From Random Search in Robots and T cells. (Conf. · 2015)
- From Microbiology to Microcontrollers: Robot search patterns inspired by T cell movement. — From Microbiology to Microcontrollers: Robot search patterns inspired by T cell movement. (Conf. · 2013)
- How Ants Turn Information into Food. — How Ants Turn Information into Food. (Conf. · 2011)
- Emergent Representation in a Robot Control Architecture. — Emergent Representation in a Robot Control Architecture. (Conf. · 2000)
- Comment on Docket No. FR-6111-P-02: HUD's Implementation of the Fair Housing Act's Disparate Impact Standard. — Comment on Docket No. FR-6111-P-02: HUD's Implementation of the Fair Housing Act's Disparate Impact Standard. (White paper)
- Community Report from the Biosignatures Standards of Evidence Workshop. — Community Report from the Biosignatures Standards of Evidence Workshop. (White paper)
- Using RuleBuilder to graphically define and visualize BioNetGen-language patterns and reaction rules. — Using RuleBuilder to graphically define and visualize BioNetGen-language patterns and reaction rules. (Chapter · 2019)
- Ant Colonies as a Model of Human Computation. — PDF (Chapter · 2014)
Development of a camera-laser-IMU system for non-contact transverse displacement estimation using an adaptive extended Kalman filter
Journal · 2026Bigger is faster in the adaptive immune response.
Journal · 2025Space-Time Causal Discovery in Earth System Science: A Local Stencil Learning Approach.
Journal · 2025CO2 emissions during the 2023 Litli Hrútur eruption in Reykjanes, Iceland: 13C tracks magma degassing.
Journal · 2024Aerial Survey Robotics in Extreme Environments: Mapping Volcanic CO2 Emissions With Flocking UAVs.
Journal · 2022The Grayness of the Origin of Life.
Journal · 2021Machine learning feature analysis illuminates disparity between E3SM climate models and observed climate change.
Journal · 2021Aerial strategies advance volcanic gas measurements at inaccessible, strongly degassing volcanoes.
Journal · 2020Quantitative Measurement of Naïve T cell Association with Dendritic Cells, FRCs, and Blood Vessels in Lymph Nodes.
Journal · 2018ROCK regulates the intermittent mode of interstitial T cell migration in inflamed lungs.
Journal · 2017Persistence and Adaptation in Immunity: T Cells Balance the Extent and Thoroughness of Search.
Journal · 2016Immune-inspired search strategies for robot swarms.
Journal · 2016Quantifying the Effect of Colony Size and Food Distribution on Harvester Ant Foraging.
Journal · 2012LoCUS: A loss-tolerant volcano survey algorithm
Conf. · 2020Brief Announcement: On Site Fidelity and the Price of Ignorance in Swarm Robotic Central Place Foraging Algorithms.
Conf. · 2019From Microbiology to Microcontrollers: Robot search patterns inspired by T cell movement.
Conf. · 2013How Ants Turn Information into Food.
Conf. · 2011Using RuleBuilder to graphically define and visualize BioNetGen-language patterns and reaction rules.
Chapter · 2019Ant Colonies as a Model of Human Computation.
Chapter · 2014UNM News Articles

UNM’s VolCAN team makes history in Canary Islands
Autonomous drones collected uncontaminated volcanic gas samples during an active eruption for carbon isotope analysis.
Read article
Engineering in Action Features Project VolCAN
UNM researchers receive an NSF National Robotics Initiative grant to develop swarms of drones for volcano research.
Read article
UNM Team Qualifies for Second Phase of NASA Robotics Challenge
Student team progress and systems work recognised as the project advances to the next phase.
Read article
Could Plants Control Robots on Mars?
A look at autonomy, sensing, and control in a space agriculture testbed.
Read article
Partnering for Success: NASA and Supercomputing Competitions
Student work across robotics and HPC showcased through national competition results.
Read article
UNM Researchers Take a Deep Dive into Our Changing Planet with SIMReef Project
Agent based reef modelling and environmental computing work highlighted through SIMReef.
Read article
CARC team gears up for SC24 Competition
Student Supercomputing Conference Competition and Student Experience at SC24.
Read article
UNM Competes in HPC Student Cluster Competitions
Training, teamwork, and results from student cluster competition participation. For more information about the teams see Ryan Scherbarth's page.
Read articleFormer Graduate Students








Former Undergraduates
It's been my priviledge to supervise outstanding undergraduate students. Getting to see them embark on their chosen careers is one of the best parts of my job.








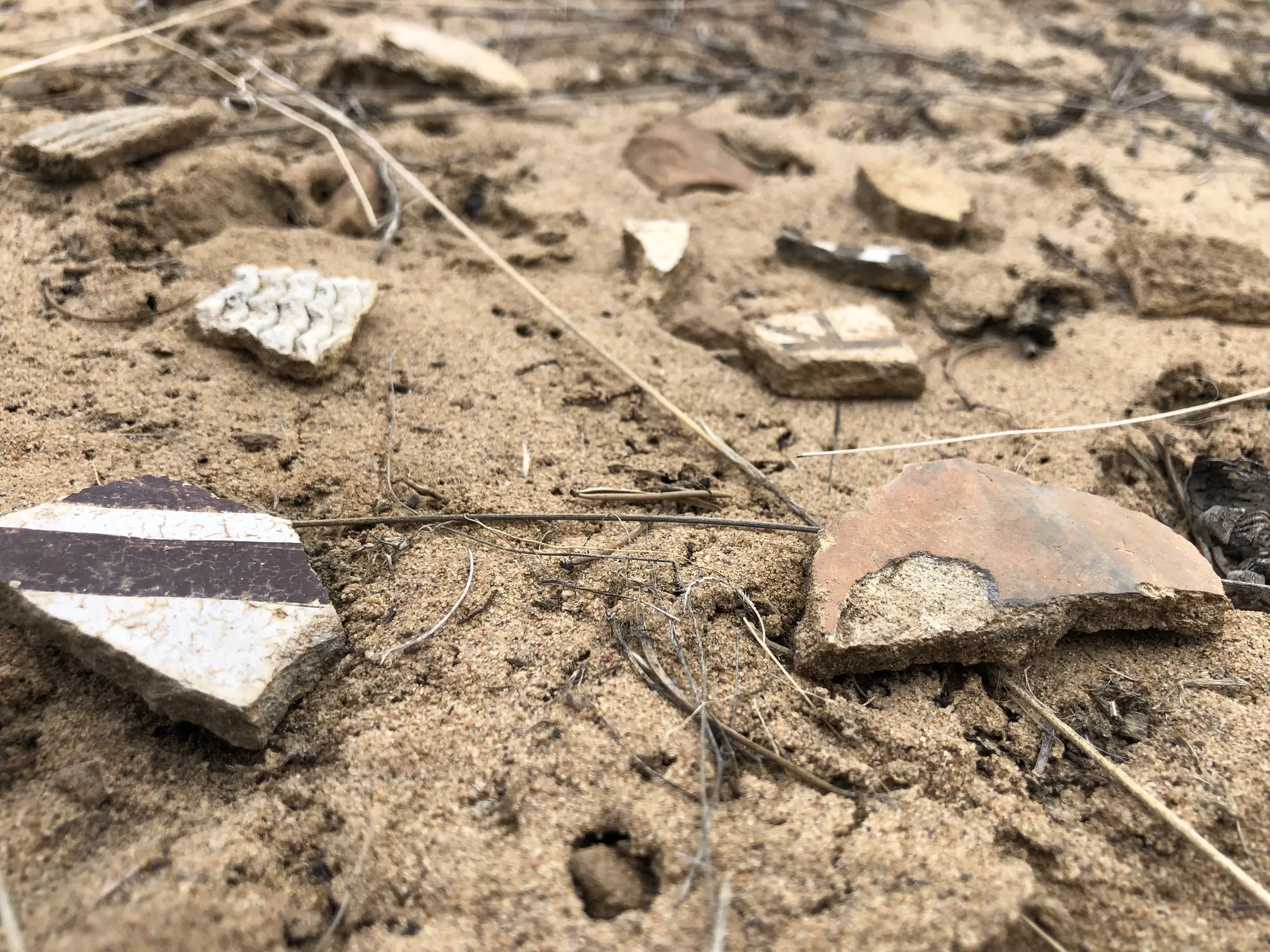
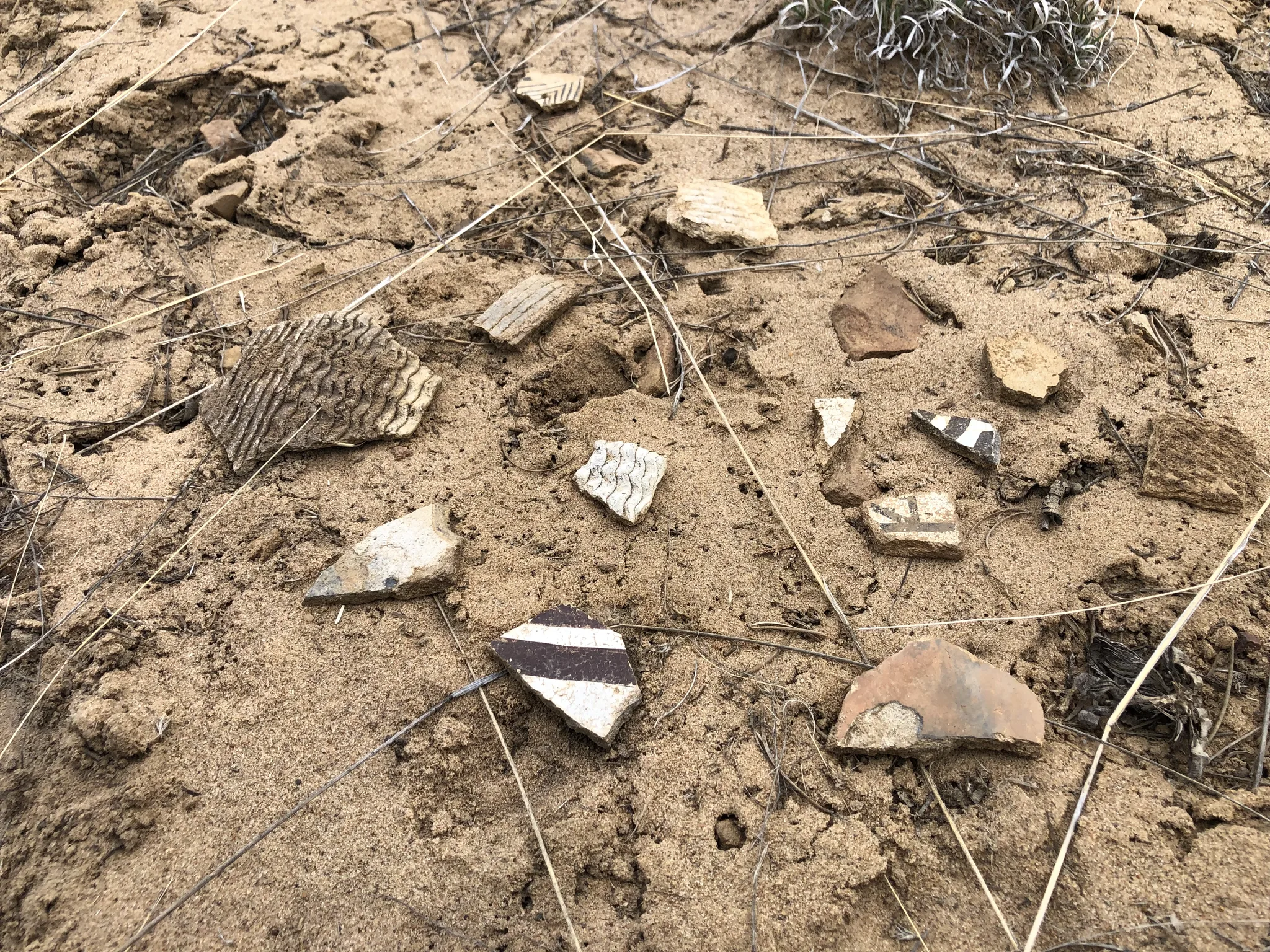
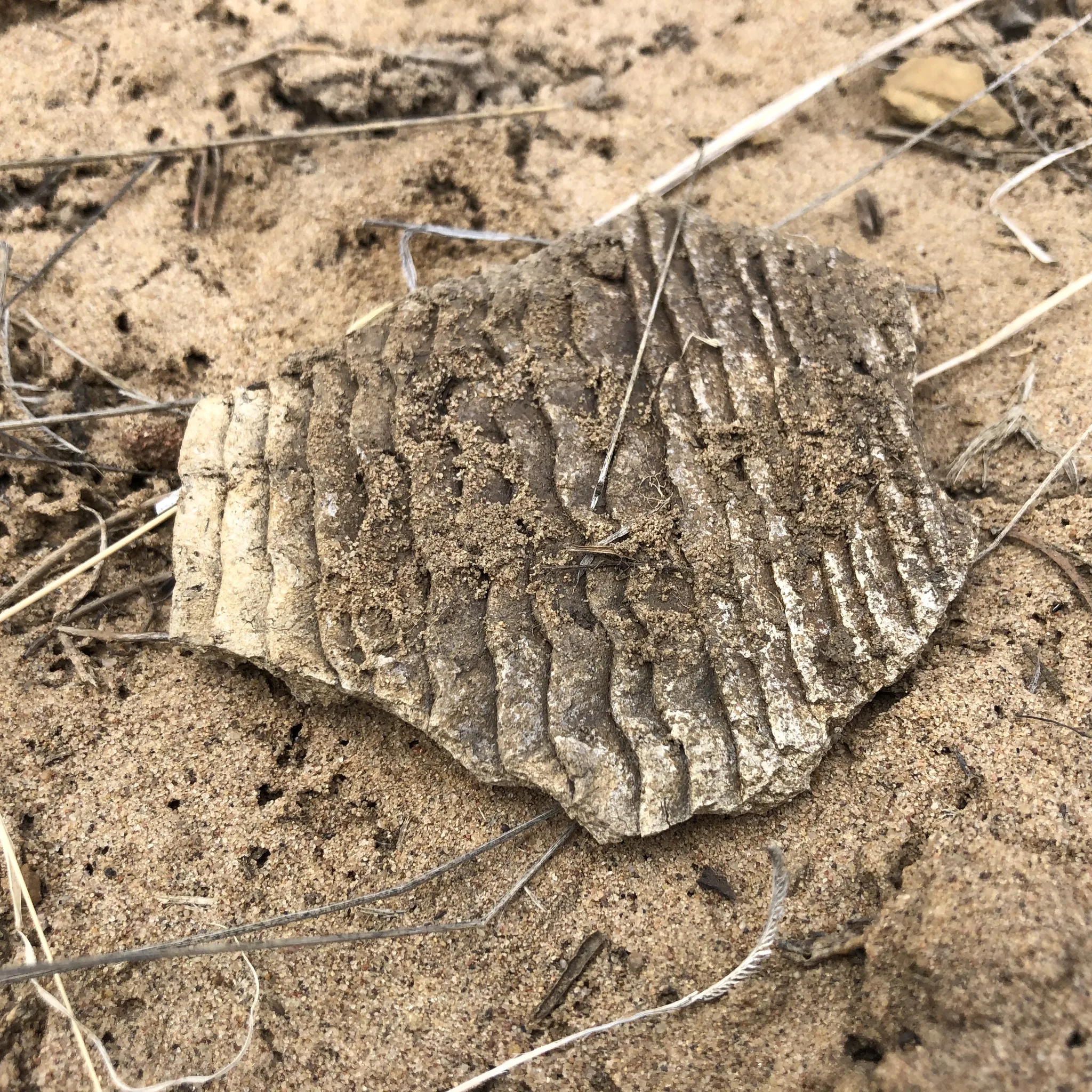
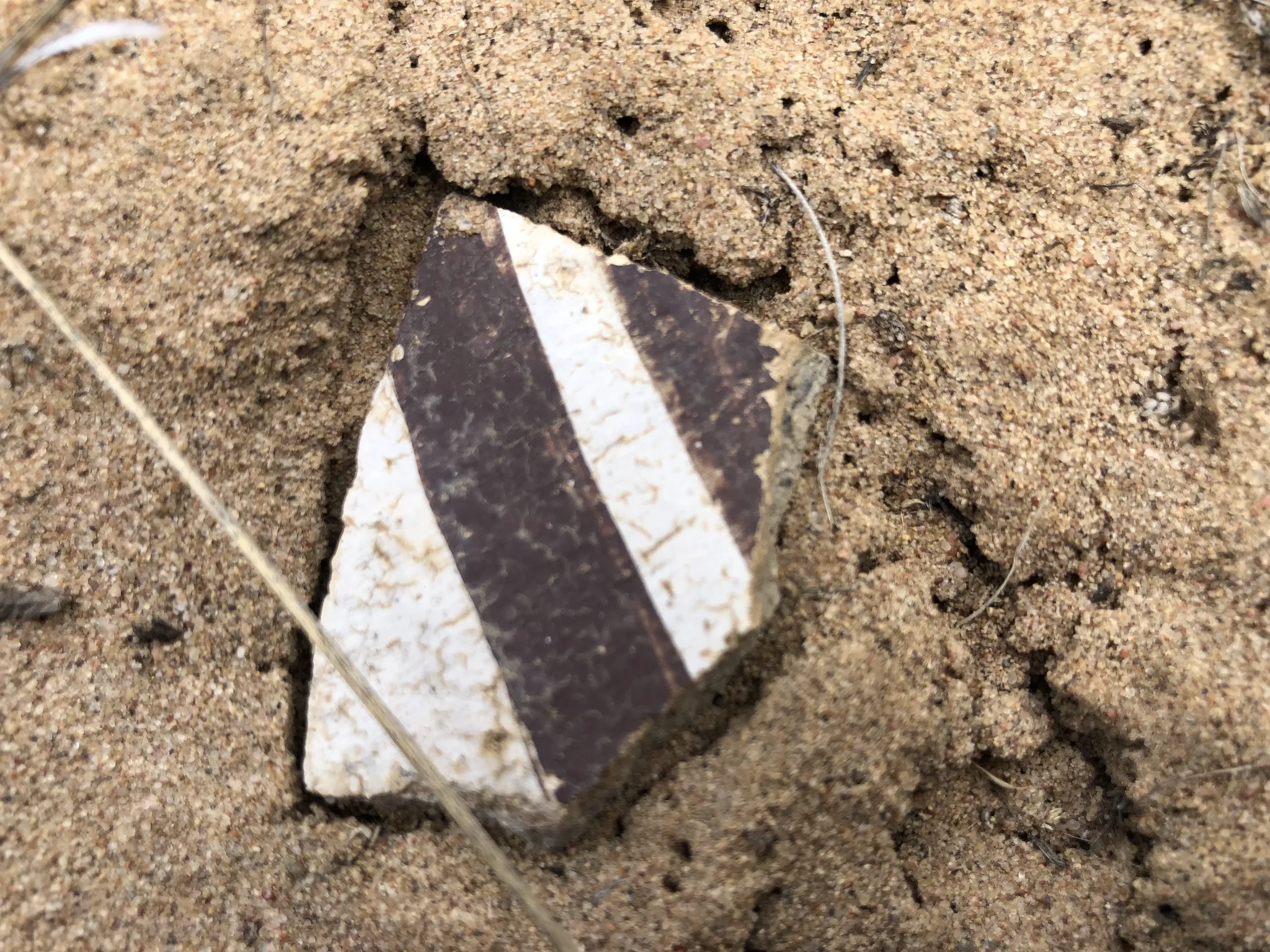
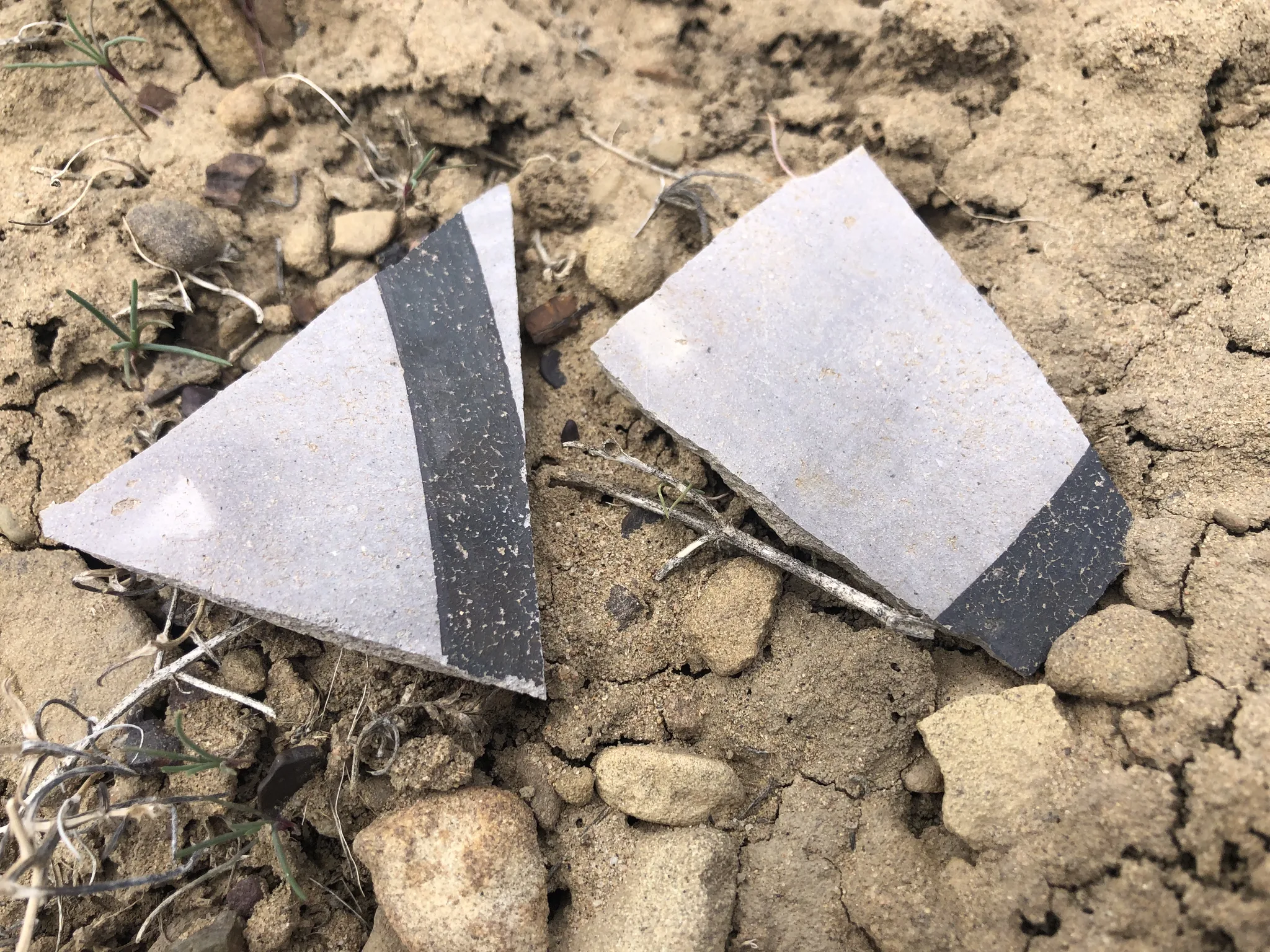
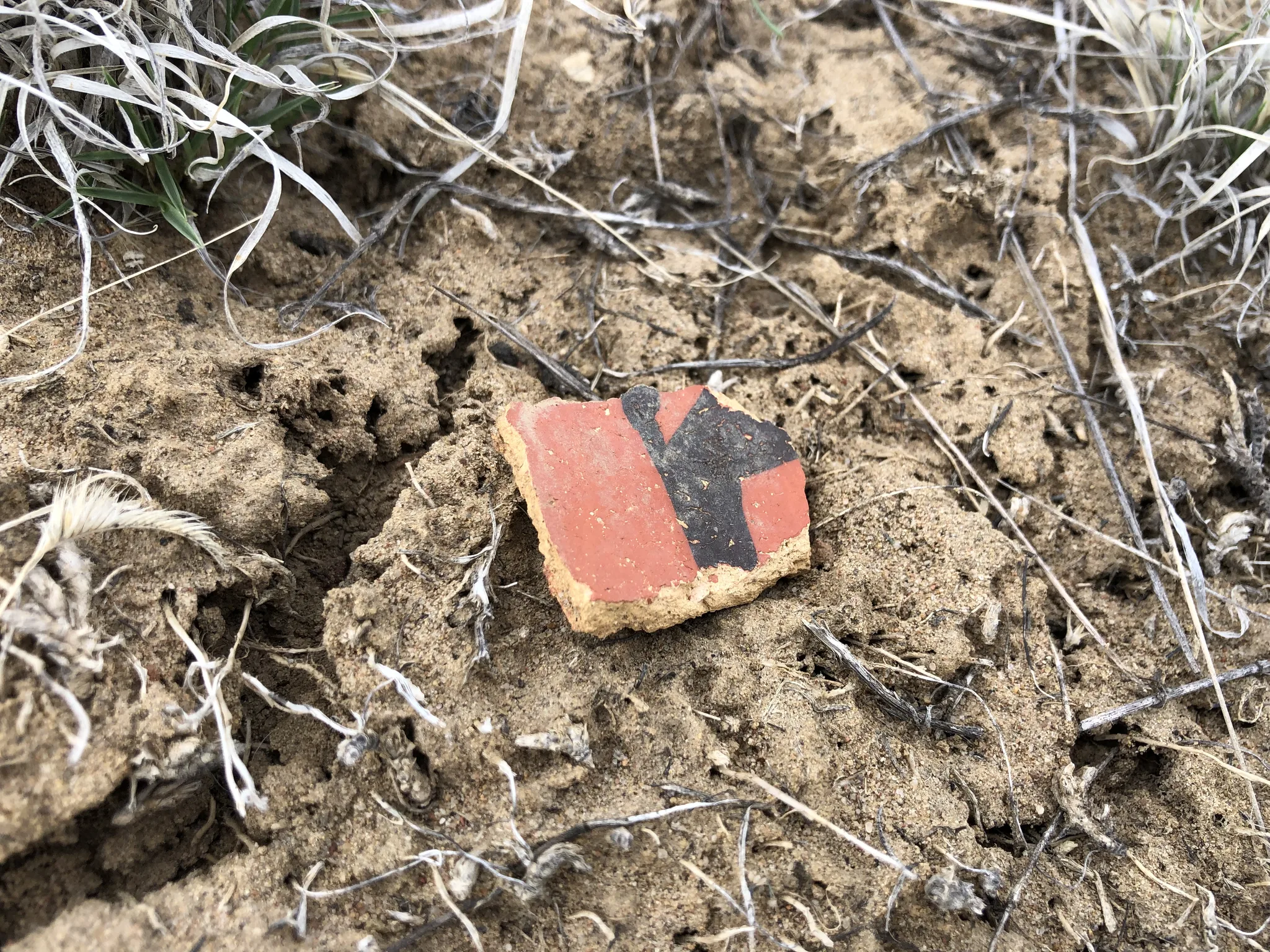
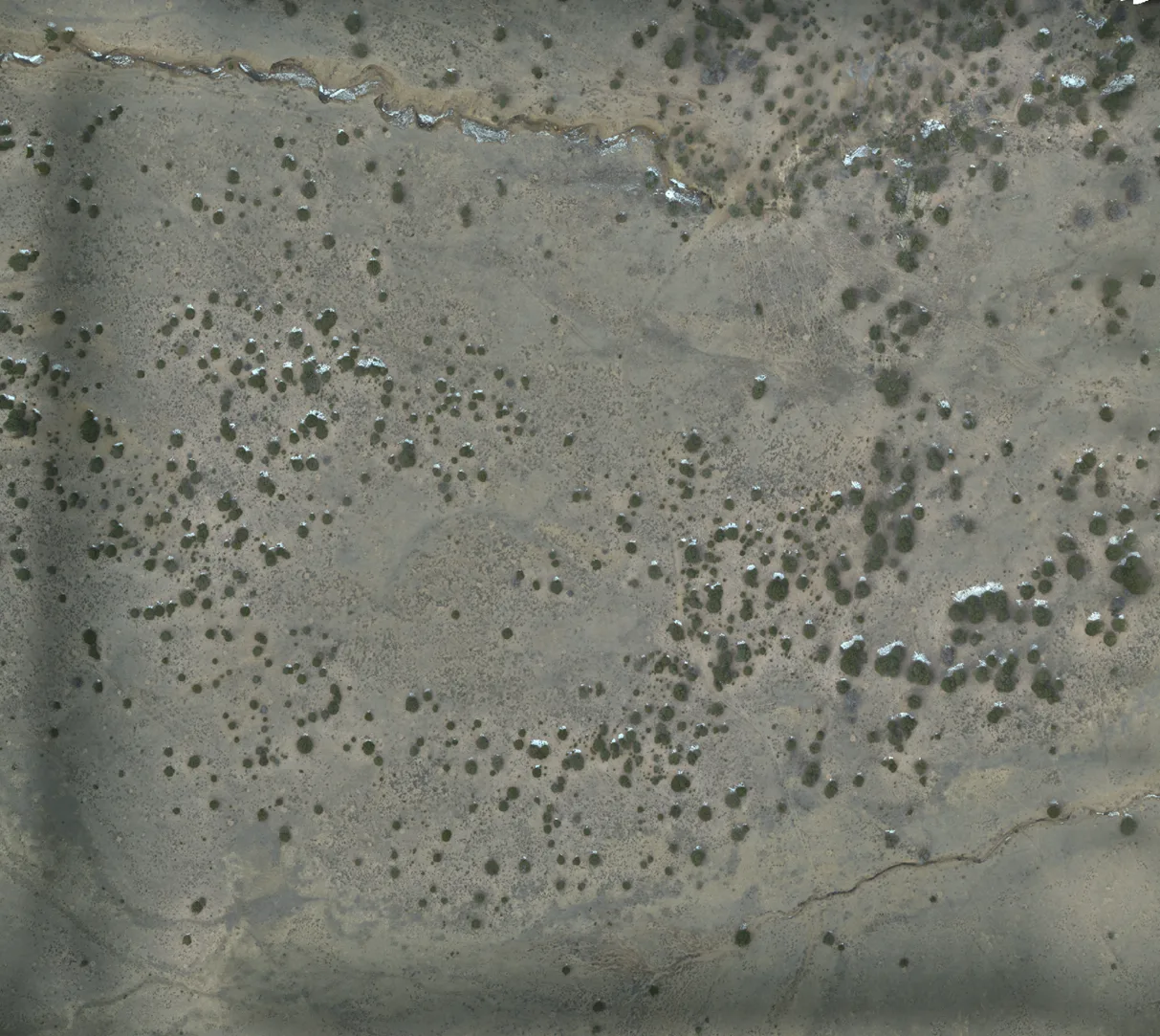
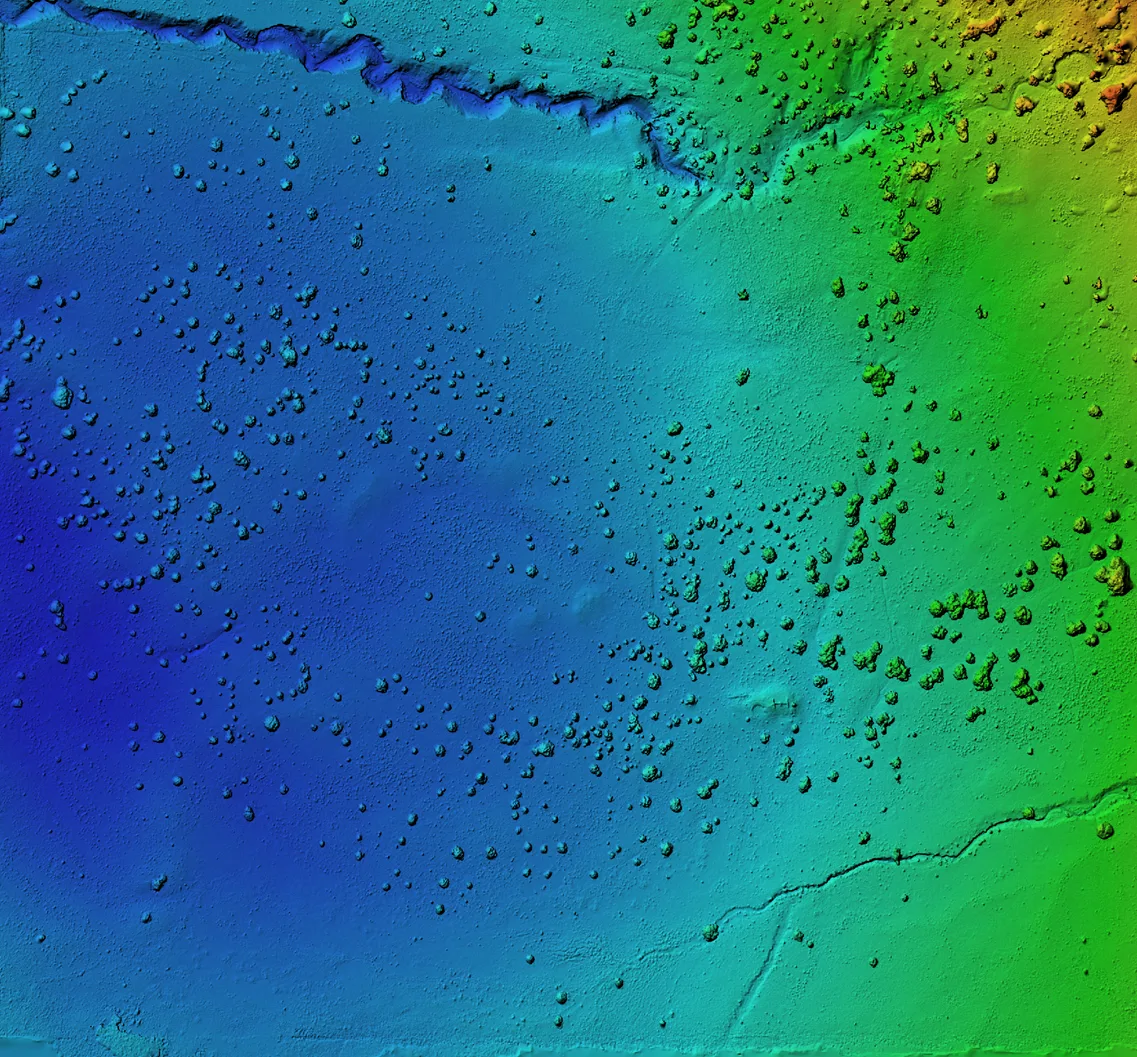
ArchaeoImaging
Dittert Site (LA 11722)
Pueblo II–III period habitation site in the Galisteo Basin, New Mexico,
characterised by surface masonry architecture, a sandstone rubble
roomblock, and a possible circular architectural feature interpreted
as a kiva. Artefact assemblage is consistent with the Pueblo II (900–1050 C.E.)
and III (1050–1300 C.E.) periods, with diagnostic examples in the ceramic
assemblages pointing to early Pueblo III.
The site is described in Dittert's
master's thesis (1949) and PhD dissertation (1959)
pp. 161–166 (pdf pp. 186–191), with discussion of the kiva on
pp. 251–255 (pdf pp. 276–280). Architectural drawings appear on
pp. 80–95 (pdf pp. 183–213).
Additional contextual information is provided by Elyea et al.,
The Armijo Canyon Archaeological Survey
,
Chapter 4 (Pueblo II/Pueblo III sites), pp. 214–216, where the site is
discussed under LA 11722 and correlated with Dittert’s L.V. 4:1–4-C.









Social Media
Facebook:
https://facebook.com/gmfricke
YouTube:
https://www.youtube.com/user/MatthewFricke
Google Scholar:
https://scholar.google.com/citations?user=hdPxTWwAAAAJ&hl=en
LinkedIn:
https://www.linkedin.com/in/gmfricke/
PGP Key
Pretty Good Privacy Email Key: http://fricke.co.uk/pgppublickey.txt
Photographs
View all photos directly in Nextcloud Memories .